| ANSI-C | 1996-1997 |

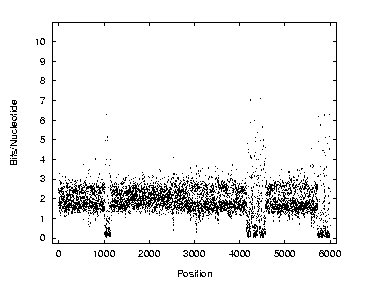
Description: This is the software used to produce the results reported in Significantly lower entropy estimates for natural DNA sequences . Its purpose is to measure the "predictability/compressibility" of a symbol sequence based on inexact matches. The local context of the position to be predicted is compared with a reference string, and all matches are noted. That is, matches of Hamming distance 0,1,2, etc. Information from each of these is combined using a novel tree-based scheme to form a prediction. The parameters of the combining tree are learned using EM (Expectation Maximization).
Our focus in Significantly lower entropy estimates for natural DNA sequences is to better understand the properties of natural DNA sequences -- not to produce an efficient software system. We are now producing an entirely new implementation of our approach to prediction/compression. This new program relies on fast inexact nearest neighbor search and online EM to produce something closer to a standard tool.